Description
We designed 100 de novo protein binders targeting human RBX1 (RING Box Protein 1), a 108-amino acid component of Cullin-RING E3 ubiquitin ligases. Our approach used BindCraft, an automated pipeline that combines AlphaFold2 backpropagation-based hallucination with ProteinMPNN sequence optimization. We ran three parallel design campaigns: (1) targeted designs directed at the E2/Glomulin-binding surface of the RING domain, informed by the crystal structure of the GLMN-RBX1-CUL1 complex (PDB: 4F52); (2) untargeted designs allowing AlphaFold2 to freely select the optimal binding site; and (3) beta-sheet-enriched designs using a negative helicity loss to generate structurally diverse binders. From 263 designs passing computational filters across all campaigns, we selected the top 100 ranked by interface pTM (i_pTM), with all designs exhibiting i_pTM >= 0.72, pLDDT >= 0.81, and sequence lengths between 60-154 amino acids.
RBX1 consists of an intrinsically disordered N-terminal region (residues 1-39) and a structured C-terminal RING-H2 finger domain (residues 40-108) stabilized by three zinc ions. We used the crystal structure of RBX1 in complex with Glomulin and CUL1 (PDB: 4F52, chain B, 3.0 A resolution) as the source for our target coordinates. The structure was trimmed to the RING domain only (residues 40-108) to minimize computational cost and focus the design on the structured, zinc-stabilized region. HETATM records including zinc ions were removed, as BindCraft operates on protein backbone coordinates; the RING fold geometry in the PDB already reflects the zinc-stabilized conformation. The chain was renamed to chain A and saved as a clean PDB file. We chose the crystal structure over the NMR ensemble (PDB: 2LGV) because crystal structures provide better-defined side chain positions at the interface, which is important for the AF2 hallucination process.
For the targeted campaign, we selected hotspot residues based on the GLMN-RBX1 interface characterized in the crystal structure (Duda et al., 2012). The interface is anchored by two complementary hydrophobic surfaces. The primary hydrophobic patch comprises RBX1 residues Ala43, Ile44, Arg46 (aliphatic portion), Trp87, Pro95, and Leu96. Secondary polar contacts include Glu55, Arg86, Arg91, and Asn98, which participate in hydrogen bonding and electrostatic interactions with GLMN. This surface is biologically validated: Glomulin binds here with ~40 nM affinity and overlaps with the E2 ubiquitin-conjugating enzyme (CDC34) binding site, as confirmed by NMR chemical shift perturbation experiments. The full hotspot specification was: A43-44, A46, A55, A86-87, A91, A95-96, A98.
All binders were generated using BindCraft (Pacesa et al., 2025), an automated de novo binder design pipeline. BindCraft uses backpropagation through the AlphaFold2 multimer network to hallucinate protein sequences predicted to bind the target, guided by a composite loss function that optimizes interface confidence (i_pTM, i_pAE), binder globularity, compactness (radius of gyration), and secondary structure content. The pipeline employs four stages: (1) soft-sequence optimization in relaxed amino acid space; (2) softmax transition to discrete sequences; (3) one-hot hard-sequence optimization; and (4) PSSM-guided semigreedy optimization. Throughout all stages, the pipeline randomly swaps between the five trained AF2 multimer models to prevent overfitting to any single model.
Hallucinated sequences are then redesigned using ProteinMPNN with soluble model weights (SolubleMPNN) to optimize core packing and surface composition while preserving the designed interface. Redesigned sequences are repredicted using the AF2 monomer model as a stringent self-consistency check. Designs passing all computational filters (pLDDT, i_pTM, i_pAE, Rosetta interface energy, shape complementarity, clash count, and binder RMSD) are accepted into the final pool.
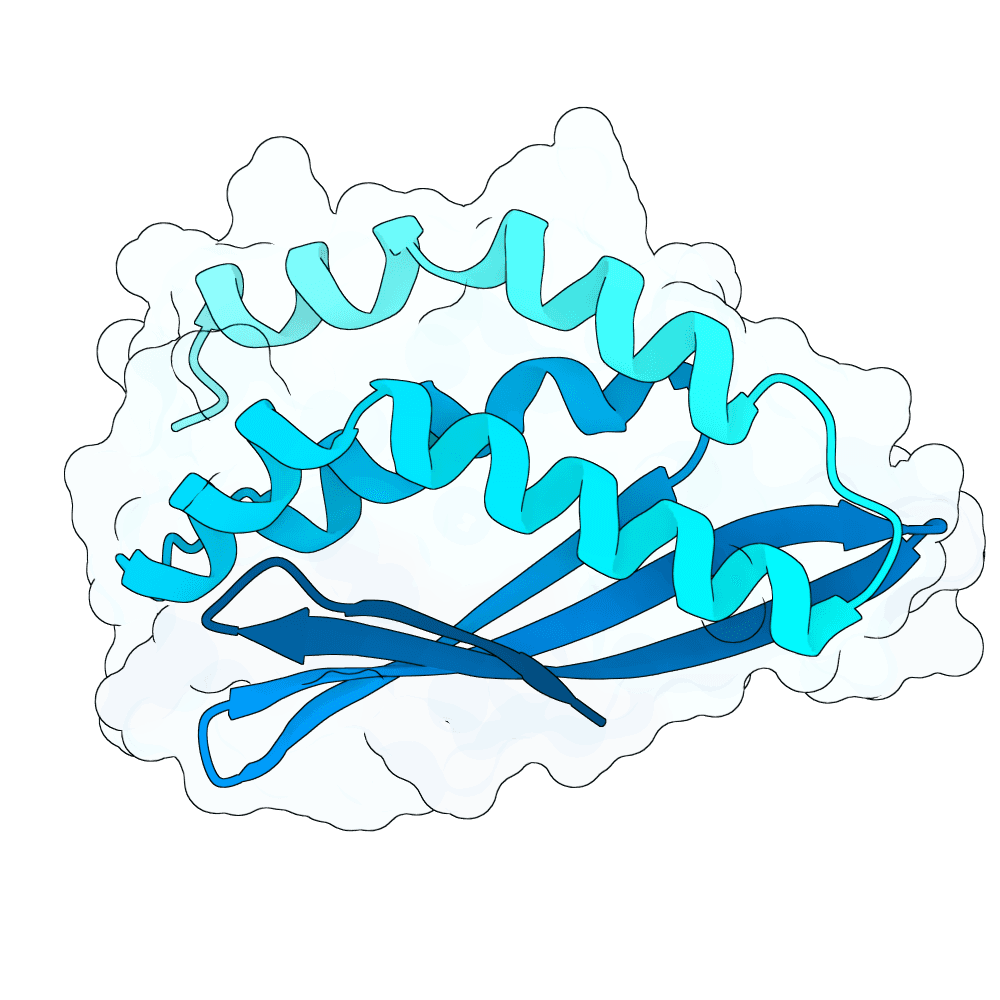
All campaigns were executed on NVIDIA A100 80GB GPUs. The targeted campaign achieved a 38% acceptance rate. From 263 total accepted designs (111 targeted, 52 free, 100 beta-sheet with relaxed filters), we selected the top 100 by average i_pTM score. The final submission comprises 66 targeted helical designs, 33 untargeted helical designs, and 1 beta-sheet design (63.6% beta-sheet content, i_pTM 0.72). All designs are fully de novo with no significant similarity to known proteins confirmed by BLAST against Swiss-Prot.
References: 1. Pacesa M, et al. (2025) Nature 646, 483-492. 2. Duda DM, et al. (2012) Mol Cell 47(3), 371-382. 3. Dauparas J, et al. (2022) Science 378, 49-56. 4. Jumper J, et al. (2021) Nature 596, 583-589.